Ligia Pereira Castro
From Laboratório de Reparo de DNA
| Revisão de 14:07, 16 Abril 2018 Root (Discussão | contribs) (→Resumo do Projeto) ← Ver a alteração anterior |
Revisão atual Root (Discussão | contribs) (→Prizes) |
||
| Linha 1: | Linha 1: | ||
| {| | {| | ||
| |- | |- | ||
| - | |[[Image:FotoLigia.jpg|150px]] | + | |[[Image:Ligia.jpg|100px]] |
| - | |Graduada em Ciências Biológicas pela Universidade Federal de São Paulo (UNIFESP). Atualmente desenvolve o projeto; Determinação de mutação genética de isolado genético de xeroderma pigmentosum em Goiás; no Instituto de Ciências Biológicas (ICB-II), Laboratório de Reparo de DNA - Prof. Dr. Carlos F.M. Menck. | + | |BSc, <b>Biological Sciences</b>, <b>Universidade Federal de São Paulo (UNIFESP)</b>.<br>PhD, <b>Biotechnology</b>, <b>Universidade de São Paulo (USP)</b>.<br><br> Has experience in Cell Biology, Molecular Biology and Next Generation Sequencing (NGS) on the identification of causative mutations for Mendelian diseases related to DNA repair deficiencies. <br><br> Ligia is a post doctoral researcher in the DNA Repair Lab - <b>Institute of Biological Sciences at University of Sao Paulo</b> since 2016. .<br> Research keywords: <b>Cancer</b>, <b>Chemotherapy</b> and <b>Rare Mendelian Disorders</b> <br> [http://buscatextual.cnpq.br/buscatextual/visualizacv.do?id=K4463877A9 Lattes]. |
| - | <br> | + | <b>Contact:</b> ligia.pereiracastro[at]gmail.com |
| - | + | ||
| |- | |- | ||
| |colspan=2| | |colspan=2| | ||
| Linha 26: | Linha 25: | ||
| <center> | <center> | ||
| - | [[Image:ArvoreLigia.png|350px]] | + | [[Image:Munfordcastroetal2.jpg|270px]] |
| + | [[Image:Munfordcastroetal.jpg|350px]] | ||
| + | [[Image:lerner.jpeg|350px]] | ||
| <br> | <br> | ||
| - | Parte da arvore genealógica da comunidade de Araras(GO) | ||
| <br> | <br> | ||
| </center> | </center> | ||
| Linha 36: | Linha 36: | ||
| Alves FIA, Moura LMS, Galante PAF, Camargo AA, Liboredo R, Pena SDJ, Sarasin A, Chaibub SC, Menck CFM | Alves FIA, Moura LMS, Galante PAF, Camargo AA, Liboredo R, Pena SDJ, Sarasin A, Chaibub SC, Menck CFM | ||
| (2017). A genetic cluster of patients with variant xeroderma pigmentosum with two different founder | (2017). A genetic cluster of patients with variant xeroderma pigmentosum with two different founder | ||
| - | mutations. British Journal of Dermatology. 176 (5), 1270-1278. <br> <br> | + | mutations. <u>British Journal of Dermatology.</u> 176 (5), 1270-1278. <br> [https://onlinelibrary.wiley.com/doi/abs/10.1111/bjd.15084 Full Text] [https://onlinelibrary.wiley.com/doi/abs/10.1111/bjd.15435 Highlighted Comment] <br><br> |
| Pedroso JL, Munford V, Bastos AU, <b>Castro LP</b>, Marussi VHR, Silva GS, Arita JH, Menck CFM, Barsottini OG | Pedroso JL, Munford V, Bastos AU, <b>Castro LP</b>, Marussi VHR, Silva GS, Arita JH, Menck CFM, Barsottini OG | ||
| - | (2017). LMNB1 mutation causes cerebellar involvement and a genome instability defect. Journal of the | + | (2017). LMNB1 mutation causes cerebellar involvement and a genome instability defect. <u>Journal of the |
| - | Neurological Sciences. 379:249-252. <br> <br> | + | Neurological Sciences.</u> 379:249-252. <br> [https://www.sciencedirect.com/science/article/pii/S0022510X1730401X?via%3Dihub Full Text] <br><br> |
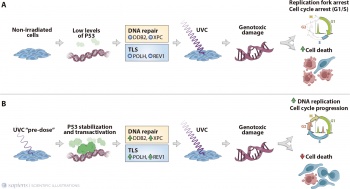
| Lerner LK, Francisco G, Soltys DT, Rocha CR, Quinet A, Vessoni AT, <b>Castro LP</b>, David TI, Bustos SO, Strauss | Lerner LK, Francisco G, Soltys DT, Rocha CR, Quinet A, Vessoni AT, <b>Castro LP</b>, David TI, Bustos SO, Strauss | ||
| BE, Gottifredi V, Stary A, Sarasin A, Chammas R, Menck CF (2016). Predominant role of DNA polymerase eta | BE, Gottifredi V, Stary A, Sarasin A, Chammas R, Menck CF (2016). Predominant role of DNA polymerase eta | ||
| - | and p53-dependent translesion synthesis in the survival of ultraviolet-irradiated human cells. Nucleic Acids | + | and p53-dependent translesion synthesis in the survival of ultraviolet-irradiated human cells. <u>Nucleic Acids |
| - | Research. 45 (3), 1270-1280.<br><br> | + | Research.</u> 45 (3), 1270-1280.<br> [https://academic.oup.com/nar/article/45/3/1270/2631187 Full Text]<br><br> |
| + | |||
| + | == Awards == | ||
| + | Tetracampeã <u> invicta </u> do Troféu Teiti Yagura (A Caixinha do Lab)<br> | ||
| + | <b> 2012 </b>: Bolo do Lázaro<br> | ||
| + | <b> 2013 </b>: Conjunto da Obra <br> | ||
| + | <b> 2016 </b>: Alan (que não fala inglês) na portaria<br> | ||
| + | <b> 2017 </b>: Garibaldi Palmisano<br> | ||
Revisão atual

| BSc, Biological Sciences, Universidade Federal de São Paulo (UNIFESP). PhD, Biotechnology, Universidade de São Paulo (USP). Has experience in Cell Biology, Molecular Biology and Next Generation Sequencing (NGS) on the identification of causative mutations for Mendelian diseases related to DNA repair deficiencies. Ligia is a post doctoral researcher in the DNA Repair Lab - Institute of Biological Sciences at University of Sao Paulo since 2016. . Research keywords: Cancer, Chemotherapy and Rare Mendelian Disorders Lattes. Contact: ligia.pereiracastro[at]gmail.com | |
|
[editar] Current Project[editar] Genotypic and phenotypic characterization of xeroderma pigmentosum (XP), cockayne syndrome (CS) and trichothiodystrophy (TTD) patients in BrazilDeficiencies in nucleotide excision repair lead to human disorders where patients
display photosensitivity and/or neurological problems, such as xeroderma pigmentosum
(XP), cockayne syndrome (CS) and trichothiodystrophy (TTD). Case reports and
genotypic descriptions of patients have been published worldwide, mainly in North
America, Europe, Africa and Japan. The follow up of these patients for decades and the study with these
cells led to the understanding of what is currently known about the molecular pathways and genetic
defects involved in the phenotypes of these syndromes. In Brazil, a few case-reports describe some patients
with these phenotypes, and genetic and molecular characterizations are scarce. With the possibility to
identify mutations directly by Next Generation Sequencing (NGS) technique, we initiated a project to
diagnose the mutations involved in these NER syndromes, mainly XP. Therefore, we also plan to investigate
the origins of these mutations and ancestries by the analysis of haplotypes with SNP-array assays.
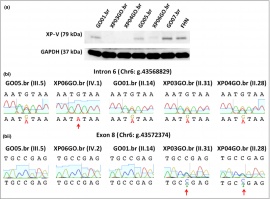
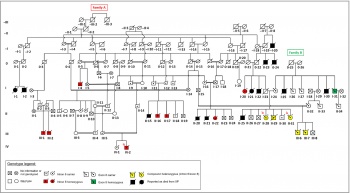
[editar] PublicationsMunford V*, Castro LP*, Souto R, Lerner LK, Vilar JB, Quayle C, Asif H, Schuch AP, de Souza TA, Ienne S,
Alves FIA, Moura LMS, Galante PAF, Camargo AA, Liboredo R, Pena SDJ, Sarasin A, Chaibub SC, Menck CFM
(2017). A genetic cluster of patients with variant xeroderma pigmentosum with two different founder
mutations. British Journal of Dermatology. 176 (5), 1270-1278. [editar] AwardsTetracampeã invicta do Troféu Teiti Yagura (A Caixinha do Lab) |
||